Targeted compound testing involves the screening of a small number of compounds using known or purported disease-modifying drugs to validate the cellular genetic disease. A number of compound library screens provide the opportunity to interrogate multiple pathways and targets in an unbiased fashion. We encourage the appreciation of all components screening in the context.
Cell-Based Drug Screening
The development of cell-based drug screening technology has enabled new opportunities to identify disease modulating therapeutics through providing an unlimited amount of pluripotent material from any patient. This process begins with the isolation of the undifferentiated cells. These cells offer the first scalable tool by providing essentially unlimited pluripotent source material. These advanced methods have greatly increased researcher access to powerful personalized medicine platforms, enabling previously unattainable in vitro disease modeling and subsequent drug screening studies. Besides, the unprecedented technology has enabled the precise study of the individual cellular components of each organ and empowered comparatively rapid drug screening for human genetic diseases.
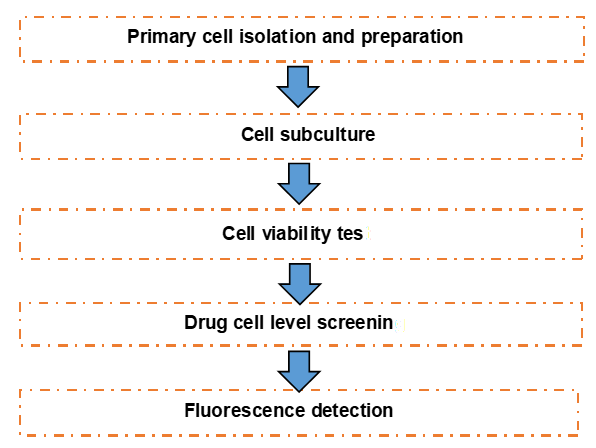 Figure 1. The workflow of cell-based drug screening.
Figure 1. The workflow of cell-based drug screening.
Virtual Screening
Virtual screening (VS) has been standard approaches in structure-based design for more than two decades. What has helped the most in drug screening is fast computing, increasing knowledge of macromolecular structures, and a substantial amount of sequence data that can be used when biological structures are lacking in drug discovery. In research, small molecules are recognized by macromolecules or interfaces eliciting a biological effect and drug discovery is based on the structure binding and its disruption to provide information on intermolecular interactions, binding conformations, potential binding pockets, and mechanistic details. With the VS, it is possible to increase the number of potential therapeutic targets and expand the focus of pharmaceuticals.
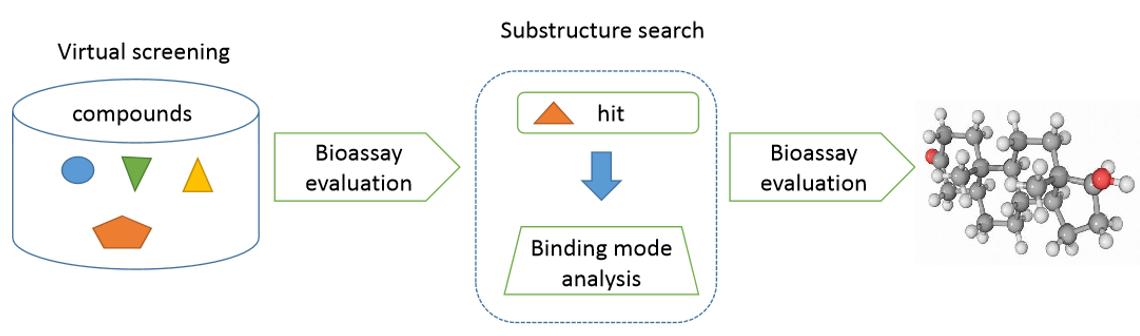 Figure 2. The diagram of virtual screening.
Figure 2. The diagram of virtual screening.
DNA Coding Library
As a powerful tool for novel drug screening, DNA-encoded-library (DEL) was first developed by Sydney Brenner of the American Scripps Institute and Richard Lerner. DEL technology is a powerful strategy to discover protein ligands. This technology involves the synthesis of a combinatorial chemical library, in which each compound is conjugated to a corresponding DNA barcode, affinity screening of the DNA-encoded library molecules against targets, and deconvolution of hit compounds by high-throughput sequencing. The DNA tags of hit beads were amplified by PCR using primers. Since this pioneering work, various DEL methods have been developed and demonstrated that several high-affinity protein ligands could be identified from DEL screening.
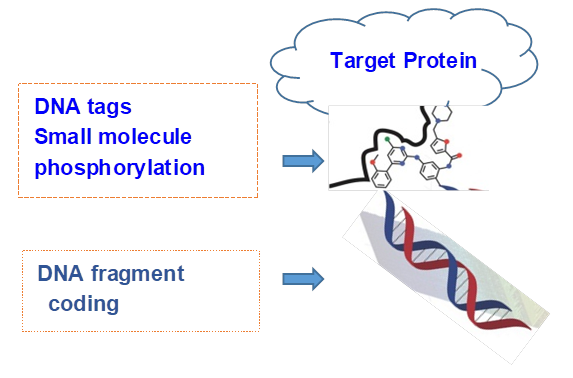 Figure 3. The general structure of a DNA-encoded technology.
Figure 3. The general structure of a DNA-encoded technology.
In the past decade, the competitive environment for the discovery of new drugs has created the need for more cost-effective and time-saving strategies to accelerate the process of identifying molecular targets in drug discovery. The successful drug discovery effort will utilize many kinds of approaches and achieve better, less expensive, and faster identification of novel leads. The technological advances observed in research will further accelerate the drug discovery process.
References
- Slater O, Kontoyianni M. The compromise of virtual screening and its impact on drug discovery. Expert Opinion on Drug Discovery, 2019, 4: 1-19.
- Brinkman R. R, et al.; Human monogenic disorders - a source of novel drug targets. Nature reviews Genetics, 2006, 7: 249-260.